Computational tools for modeling and data analyses (PIs: P. Bernadó & N. Sibille)
People Involved: P. Bernadó, N. Sibille, F. Allemand, F. Herranz-Trillo, A. Estana, A. Sagar
During the present contract the group has developed an intense activity in developing new tools for the structural modeling of flexible systems as well as the comprehensive analysis of complex SAXS data. In the following paragraphs the most relevant results on these two research lines will be described.
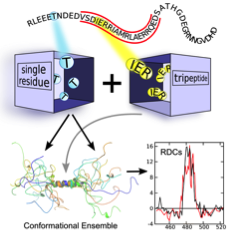
- Structural modeling of Disordered Proteins: Structural modeling of IDPs is extremely challenging as their special features are coded in the conformational bias at the residue-level. In order to solve of these challenges, we have developed a tripeptide database from high-resolution structures in collaboration with Juan Cortés that allows to build realistic IDP ensemble models for these proteins (Estaña et al, 2019). The resulting ensembles are in agreement with extensive experimental NMR and SAXS data. The enhanced performance of this approach is based on the inclusion of the specific chemical and structural properties of the neighboring residues. The tripeptide database is extremely rich and can also be used to decipher the folding pathway of small structural elements of proteins (Estaña et al., 2019) and the prediction of partially structured elements in IDPs from their sequences (Estaña et al., in preparation).
- Chemometric analysis of SAXS data from complex biomolecular systems: Many biological systems present distinct species that are in equilibrium. These polydisperse systems are extremely difficult to characterize as the experimental observables are population-weighted averages. SAXS is very sensitive to the overall size and shape of particles and, therefore, it can monitor changes when the equilibrium is perturbed by modifying the experimental conditions (concentration, pH, time...). We have developed a chemometric approach, which we call COSMiCS, to decompose large population-weighted SAXS datasets. COSMiCS uses multiple SAXS data representations to decrease the degeneracy of possible solutions. As a proof of concept, we applied COSMiCS to study the fibrillation of insulin and α-synuclein by SAXS for which the population and structure of the three main species (monomer, oligomer and fibril) in equilibrium were derived (Herranz-Trillo et al., 2017).
Main collaborators: J. Cortés (LAAS, Toulouse), B. Vestergaard (U. Copenhagen), R. Tauler (CSIC, Bercelona)