We explore the structure and dynamics of membrane assemblies, including mechanical properties, membrane proteins structure and interaction, and membrane remodeling mechanisms, on living cells or artificial membrane models, using single molecule experimental approaches such as atomic force (AFM) and fluorescence/super resolution microscopies.
Remodeling and partition of membrane components
L. Costa, P.E. Milhiet, C. Bénistant, P. Dosset, C. Doucet, C. Godefroy, L. Fernandez
We are interested in the molecular mechanisms of lipid-protein interaction within membranes and their spatiotemporal recruitment in different contexts, especially cancer and infection.
Structure and dynamics of tetraspanins. Tetraspanins are transmembrane proteins forming a network of protein-protein-lipid interactions at the plasma membrane of eukaryotic cells. They are involved in numerous cell functions and associated with several diseases. Using single molecule fluorescence microscopy and AFM we are exploring tetraspanin lateral segregation into micro- or nanodomains. We are especially investigating the dynamics and partitioning of the metastasis suppressor CD82 and its rela tionship to cell migration, and the role of tetraspanins CD9 and CD81 in cell infection by HIV-1 and Influenza viruses.
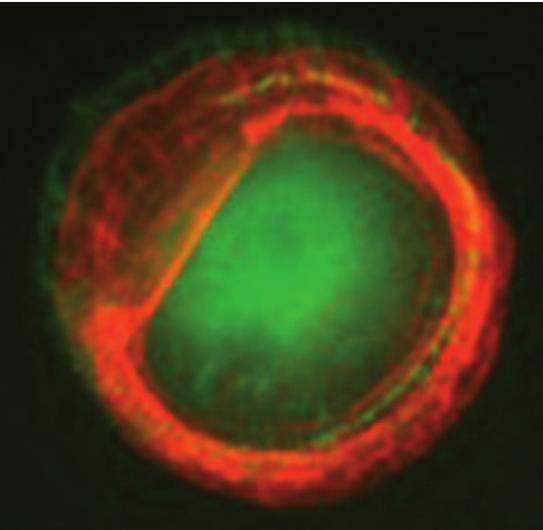
Actin and paxillin labeling of micropatterned cells expressing CD82
Septin organization during membrane remodeling. Septins are conserved GTP-binding proteins involved in membrane compartmentalization and remodeling. Their ability to form functional filaments depends on their binding to PIP2 lipids. Using supported lipid bilayers observed with high-speed AFM and correlative AFM-fluorescence microscopy, we explore at the meso and nanoscale how these proteins polymerize, recruit lipids and deform biological membranes.
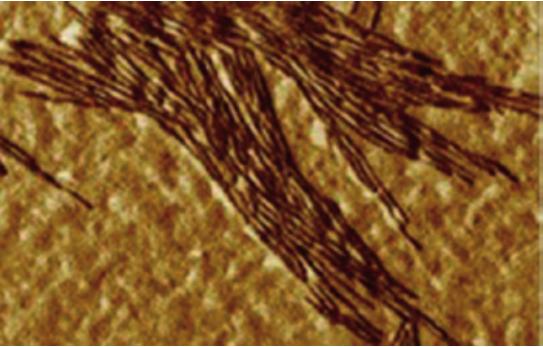
AFM micrograph of yeast septin imaged in air
Collaborations: E. Rubinstein (Villejuif), D. Levy (Institut Curie), F. Berditchevski (Birmingham, Royaume-Uni), O. Piétrement (Institut Gustave Roussy), G. Drin (IPMC, Nice), M. Mougel (IRIM, Montpellier), Sanofi-Pasteur.
Nanomechanics of cancer metastasis
C. Bénistant, L. Costa, C. Godefroy, P.E. Milhiet
Physical properties of the cancer cell microenvironment play important roles in promoting cell invasive migration phenotypes and can constitute an obstacle for therapy accessibility. We aim at analyzing how biomechanics impact the formation and progression of cancer by measuring physical properties of cancer cells and tissues, using AFM (force spectroscopy and force mapping).
Mechanical properties of colorectal cancers in transition to metastasis. We measure and correlate the rigidity (Young Modulus) of a bank of colorectal cancer tissues with genomic and patient's survival data, in collaboration with clinicians of the Montpellier Cancer Research Center.
CD82 metastasis suppressor functions. CD82 regulates several membrane remodeling events such as adhesion, migration, endocytosis... by unknown mechanisms. As membrane organizers, tetraspanins could influence membrane nanomechanics. We test the influence of CD82 on membrane tension, a key parameter in membrane remodeling.
Collaborations: E. Crapez and A. Turtoi (IRCM, Montpellier)
Mechanisms of nuclear pore complex assembly
C. Doucet, L. Costa, P. Dosset, P.E. Milhiet, A. Vial
Nuclear Pore Complexes (NPCs) are the only gateways between the nucleus and cytoplasm. In dividing cells, the number of NPCs doubles during interphase. New pore insertion in the nuclear envelope requires the local fusion of inner and outer nuclear membranes. This implies exquisite coordination between membrane remodeling and nucleoporins recruitment and assem bly. We aim at understanding the mechanisms involved in early steps of interphase NPC formation, in particular nuclear membrane deformation, following 2 main axes:
1. Identify the sequential involvement of molecular players in membrane deformation. We combine AFM and Single Molecule Localization Microscopy on isolated nuclei, to correlate the recruitment of specific proteins with nuclear membrane deformation and pore assembly.
2. Understand the peculiar deformation of the nuclear envelope. Membrane bending agents suspected to intervene in pore formation induce convex deformations on synthetic liposomes while pore intermediates are concave. We investigate determinants of the pore membrane leading to this discrepancy.
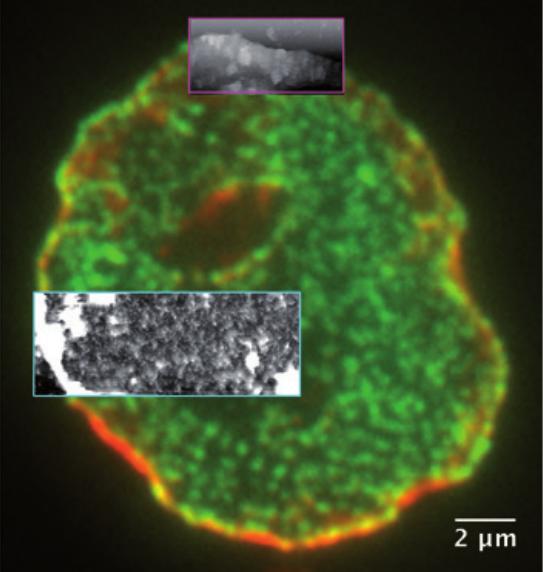
Correlative AFM and fluorescence microscopy
Structural dynamics of single metabotropic glutamate receptors
E. Margeat, C. Clerté, R. Quast
Using a combination of advanced single molecule microscopy methods, we investigate the activation mechanism of G-protein coupled receptors.
Metabotropic glutamate receptors (mGluR) are members of the class C G-protein coupled Receptors (GPCR family). Each subunit is composed of an extracellular domain that binds orthosteric ligands such as glutamate, and a heptahelical transmembrane domain responsible for G-protein activation. We study mGluR structural dynamics by a combination of site-specific labeling, receptor purification and solubilization, and smFRET with multiparametric fluorescence detection (MFD) .
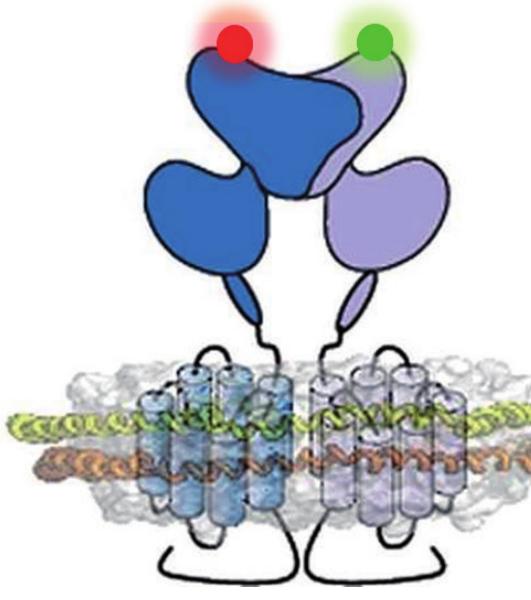
Cartoon representation of a metabotropic glutamate receptor dimer in a lipid bilayer
Collaborations: J.P. Pin (IGF, Montpellier), C. Seidel (Düsseldorf, Allemagne), D. Levy, M. Dahan (Institut Curie, Paris)
L. Costa, E. Margeat, P.E. Milhiet, C. Clerté
Our research requires the development of new methodological approaches in the fields of fluorescence spectroscopy, microscopy and atomic force microscopy, such as:
- Correlative Atomic Force with super-resolution fluorescence microscopy (STORM and Light Sheet Microscopy (LSM))
- Non-contact, photonic-based near-field probe microscopy
- One- and two-photon scanning fluorescence fluctuation microscopies
- Two-spot correlation spectroscopy coupled with rheometry
- Photonic structures for high resolution single molecule FRET
Collaborations: M. Nollmann, M. Abkarian (CBS), T. Ando (Kanazawa, Japon), C. Seidel (Düsseldorf, Allemagne), J. Wenger (Institut Fresnel, Marseille), E. Lesniewska (Institut Carnot, Dijon)

